Lipidome Juxtaposition (LUX score) Error Modeling
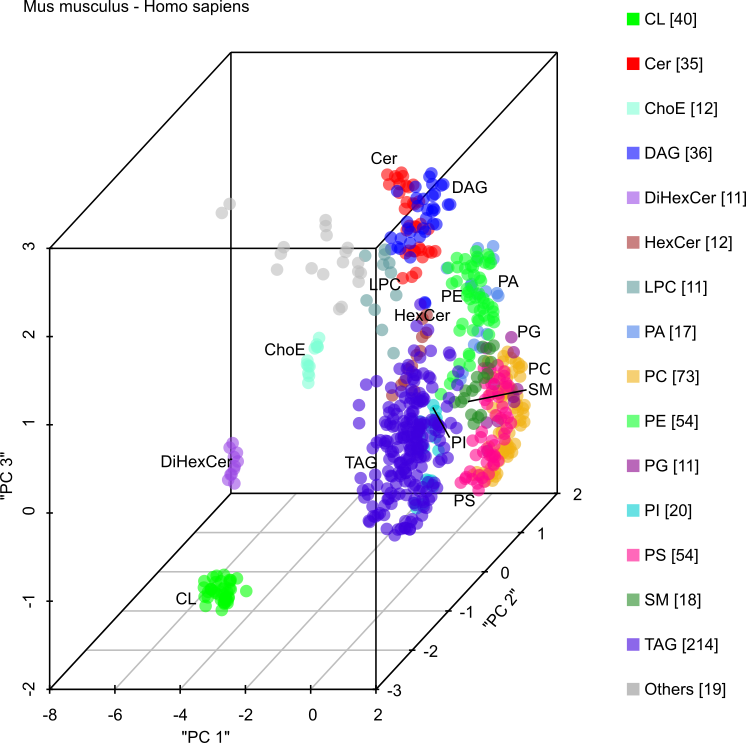
LUX score is the first metric for comparing lipidomes. It provides a measure of homology between lipidomes similar to genetic analyses (Marella et al. 2015, PLOS Comp. Biol).
This application provides a form-based, web-based graphical user inteface to run error modeling for LUX score.
The error model can be used to modify the presence and quantity low abundant lipids to estimate the robustness for the LUX score clustering. For example, it can show if the common lipid count does comprise a sufficiently robust tree topology and groups lipidomes systematically different. Additionally, applying such an experiment may show you that the compositional differences itself are useful to assign a phenotype while comparison purely based on quantities are dominated by changes of abundant lipids.
An error model for lipidome analysis is computed by iterating over all lipid quantities x of the original data set according to the following algorithm:
for each lipid with abundance x
x' = x + rnorm(1,μ,σ)
if x' > threshold
return x'
The aim is to count the number of bipartitions found in a series of lipidomes after N iterations of the algorithm.
- Dominik Schwudke, Andrej Shevchenko, Nils Hoffmann and Robert Ahrends, Lipidomics Informatics for Life-Science, Journal of Biotechnology, J Biotechnol. 261:131-136, 2017.
- Lars F. Eggers, Julia Müller, Chakravarthy Marella, Verena Scholz, Henrik Watz, Christian Kugler, Klaus F. Rabe, Torsten Goldmann & Dominik Schwudke, Lipidomes of lung cancer and tumour-free lung tissues reveal distinct molecular signatures for cancer differentiation, age, inflammation, and pulmonary emphysema, Scientific Reports, Sci Rep. 7(1):11087, 2017.
- Chakravarthy Marella, Andrew E. Torda and Dominik Schwudke, The LUX score: a metric for lipidome homology, PLoS computational biology, PLoS Comput Biol. 11(9):e1004511, 2015.



